Scientific research
Current Location:Home 〉Research
- Liang, Y.; Jiao, S.; Wang, M.; Yu, H.; Shen, Z., A CRISPR/Cas9-based genome editing system for Rhodococcus ruber TH. Metabolic Engineering 2019. DOI: 10.1016/j.ymben.2019.10.003.
- Cheng, F. Y.; Yu, H. M.; Stephanopoulos, G., Engineering Corynebacterium glutamicum for high-titer biosynthesis of hyaluronic acid. Metabolic Engineering 2019, 55, 276-289.
- Wang, M.; Qi, W.; Xu, H.; Yu, H.; Zhang, S.; Shen, Z., Affinity-binding immobilization of D-amino acid oxidase on mesoporous silica by a silica-specific peptide. Journal of industrial microbiology & biotechnology 2019.
- Tang, L.; Yang, J.; Chen, J.; Zhang, J.; Yu, H.; Shen, Z., Design of salt-bridge cyclization peptide tags for stability and activity enhancement of enzymes. Process Biochemistry 2019, 81, 39-47.
- Cheng, F.; Luozhong, S.; Yu, H.; Guo, Z., Biosynthesis of Chondroitin in Engineered Corynebacterium glutamicum. Journal of Microbiology and Biotechnology 2019, 29, (3), 392-400.
- Wang, M.; Yu, H.; Shen, Z., Antisense RNA-Based Strategy for Enhancing Surfactin Production in Bacillus subtilis TS1726 via Overexpression of the Unconventional Biotin Carboxylase II To Enhance ACCase Activity. Acs Synthetic Biology 2019, 8, (2), 251-256.
- Liang, Y.; Jiao, S.; Wang, M.; Yu, H.; Shen, Z., Overexpression of epoxide hydrolase in Rhodococcus ruber with high robustness for the synthesis of chiral epichlorohydrin. Process Biochemistry 2019, 79, 49-56.
- Guo, M.; Yang, L.; Li, J.; Jiao, S.; Wang, Y.; Luo, G.; Yu, H., Effects of interface adsorption of Rhodococcus ruber TH3 cells on the biocatalytic hydration of acrylonitrile to acrylamide. Bioprocess and Biosystems Engineering 2018, 41, (7), 931-938.
- Wang, M.; Chen, J.; Yu, H.; Shen, Z., Improving stress tolerance and cell integrity of Rhodococcus ruber by overexpressing small-shock-protein Hsp16 of Rhodococcus. Journal of Industrial Microbiology & Biotechnology 2018, 45, (10), 929-938.
- Chen, Y.; Jiao, S.; Wang, M.; Chen, J.; Yu, H., A novel molecular chaperone GroEL2 from Rhodococcus ruber and its fusion chimera with nitrile hydratase for co-enhanced activity and stability. Chemical Engineering Science 2018, 192, 235-243.
- Jiao, S.; Yu, H.; Shen, Z., Core element characterization of Rhodococcus promoters and development of a promoter-RBS mini-pool with different activity levels for efficient gene expression. New Biotechnology 2018, 44, 41-49.
- Wang, Q.; Yu, H.; Wang, M.; Yang, H.; Shen, Z., Enhanced biosynthesis and characterization of surfactin isoforms with engineered Bacillus subtilis through promoter replacement and Vitreoscilla hemoglobin co-expression. Process Biochemistry 2018, 70, 36-44.
- Cheng, F.; Luozhong, S.; Guo, Z.; Yu, H.; Stephanopoulos, G. J. B. J., Enhanced biosynthesis of hyaluronic acid using engineered Corynebacterium glutamicum via metabolic pathway regulation. 2017, 12, (10), 1700268.
- Yang, J.; Cheng, F.; Yu, H.; Wang, J.; Guo, Z.; Stephanopoulos, G., Key Role of the Carboxyl Terminus of Hyaluronan Synthase in Processive Synthesis and Size Control of Hyaluronic Acid Polymers. Biomacromolecules 2017, 18, (4), 1064-1073.
- Jiao, S.; Chen, J.; Yu, H.; Shen, Z., Tuning and elucidation of the colony dimorphism in Rhodococcus ruber associated with cell flocculation in large scale fermentation. Applied Microbiology and Biotechnology 2017, 101, (16), 6321-6332.
- Zhang, S.; Yu, H.; Yang, J.; Shen, Z., Design of the nanoarray pattern Fe-Nibi-metal nanoparticles@M13 virus for the enhanced reduction of p-chloronitrobenzene through the micro-electrolysis effect. Environmental Science-Nano 2017, 4, (4), 876-885.
- Li, J.; Guo, M.; Jiao, S.; Wang, Y.; Luo, G.; Yu, H., A kinetic study of the biological catalytic hydration of acrylonitrile to acrylamide. Chemical Engineering Journal 2017, 317, 699-706.
- Jiao, S.; Li, X.; Yu, H.; Yang, H.; Li, X.; Shen, Z., In situ enhancement of surfactin biosynthesis in Bacillus subtilis using novel artificial inducible promoters. Biotechnology and Bioengineering 2017, 114, (4), 832-842.
- Luo, H.; Zhu, L.; Chang, Y.; Liu, X.; Liu, Z.; Sun, H.; Li, X.; Yu, H.; Shen, Z., Microenvironmental pH changes in immobilized cephalosporin C acylase during a proton-producing reaction and regulation by a two-stage catalytic process. Bioresource Technology 2017, 223, 157-165.
- Sun, J.; Yu, H.; Chen, J.; Luo, H.; Shen, Z., Ammonium acrylate biomanufacturing by an engineered Rhodococcus ruber with nitrilase overexpression and double-knockout of nitrile hydratase and amidase. Journal of Industrial Microbiology & Biotechnology 2016, 43, (12), 1631-1639.
- Li, J.; Chen, Y.; Guo, M.; Wang, Y.; Xu, J.; Luo, G.; Yu, H., Using microchannels to visually investigate the formation and dissolution of acrylonitrile droplets in a bio-hydration system. Chemical Engineering Science 2016, 152, 239-247.
- Wei, Y.; Luo, H.; Chang, Y.; Yu, H.; Shen, Z., Reversible immobilization of cephalosporin C acylase on epoxy supports coated with polyethyleneimine. Biocatalysis and Biotransformation 2015, 33, (5-6), 250-259.
- Cheng, F.; Gong, Q.; Yu, H.; Stephanopoulos, G., High-titer biosynthesis of hyaluronic acid by recombinant Corynebacterium glutamicum. Biotechnology Journal 2016, 11, (4), 574-584.
- Tian, Y.; Chen, J.; Yu, H.; Shen, Z. J. J. M. B., Overproduction of the Escherichia coli chaperones GroEL-GroES in Rhodococcus ruber improves the activity and stability of cell catalysts harboring a nitrile hydratase. 2015, 26, (2), 337.
- Zhang, S.; Nakano, K.; Zhang, S.-l.; Yu, H.-m., Synthesis of dispersive iron or iron-silver nanoparticles on engineered capsid pVIII of M13 virus with electronegative terminal peptides. Journal of Nanoparticle Research 2015, 17, (10).
- Luo, H.; Ma, J.; Chang, Y.; Yu, H.; Shen, Z., Directed Evolution and Mutant Characterization of Nitrilase from Rhodococcus rhodochrous tg1-A6. Applied Biochemistry and Biotechnology 2016, 178, (8), 1510-1521.
- Yang, H.; Yu, H.; Shen, Z., A novel high-throughput and quantitative method based on visible color shifts for screening Bacillus subtilis THY-15 for surfactin production. Journal of Industrial Microbiology & Biotechnology 2015, 42, (8), 1139-1147.
- Huan, Y.; Xu, L.; Xue, L.; Huimin, Y.; Zhongyao, S. J. A.; Chemistry, B., Identification of lipopeptide isoforms by MALDI-TOF-MS/MS based on the simultaneous purification of iturin, fengycin, and surfactin by RP-HPLC. 2015, 407, (9), 2529-2542.
- Li, X.; Yang, H.; Zhang, D.; Li, X.; Yu, H.; Shen, Z., Overexpression of specific proton motive force-dependent transporters facilitate the export of surfactin in Bacillus subtilis. Journal of Industrial Microbiology & Biotechnology 2015, 42, (1), 93-103.
- He, H.; Wei, Y.; Luo, H.; Li, X.; Wang, X.; Liang, C.; Chang, Y.; Yu, H.; Shen, Z., Immobilization and Stabilization of Cephalosporin C Acylase on Aminated Support by Crosslinking with Glutaraldehyde and Further Modifying with Aminated Macromolecules. Biotechnology Progress 2015, 31, (2), 387-395.
- Luo, H.; Zhao, H.; Chang, Y.; Wang, Q.; Yu, H.; Shen, Z., Oriented Immobilization and Characterization of a Poly-Lysine-Tagged Cephalosporin C Acylase on Glyoxyl Agarose Support. Applied Biochemistry and Biotechnology 2015, 175, (4), 2114-2123.
- Li, J.; Liu, J.; Chen, J.; Wang, Y.; Luo, G.; Yu, H., Multiple reuses of Rhodococcus ruber TH3 free cells to produce acrylamide in a membrane dispersion microreactor. Bioresource Technology 2015, 187, 198-204.
- Li, J.; Yang, L.; Ding, X.; Chen, J.; Wang, Y.; Luo, G.; Yu, H., Visual study of mass transfer characterization in the process of biological catalytic hydration of acrylonitrile using pendant drop method. Rsc Advances 2015, 5, (96), 79164-79171.
- Li, J.; Chen, J.; Wang, Y.; Luo, G.; Yu, H., Hydration of acrylonitrile to produce acrylamide using biocatalyst in a membrane dispersion microreactor. Bioresource Technology 2014, 169, 416-420.
- Chen, J.; Jia, X.; Yu, H.; Luo, H.; Shen, Z., Catalytic kinetics of recombinant nitrile hydratase coupling terminal salt bridge and point-mutation. CIESC Journal 2014, 65, (7), 2821-2828.
- Zhang, J.; Yu, H.; Wang, Y.; Luo, H.; Shen, Z. J. J. o. B., Determination of the second autoproteolytic cleavage site of cephalosporin C acylase and the effect of deleting its flanking residues in the α-C-terminal region. 2014, 184, (6), 138-145.
- Shuai, Z.; Nakano, K.; Yu, H.; Shen, Z. J. M. R. E., Bio-nano complexes of ZVFeNPs/Fe-s-M13 and Cd (II)/Cd-s-M13 accelerate Cd (II) reduction by FeNPs through dual dispersing and separate deposition. 2014, 1, (1), 015043.
- Chen, J.; Yu, H.; Liu, C.; Liu, J.; Shen, Z., Improving stability of nitrile hydratase by bridging the salt-bridges in specific thermal-sensitive regions. Journal of Biotechnology 2013, 164, (2), 354-362.
- Zhang, J.; Wang, Z.-T.; Yu, H.-M.; Ma, Y., Paenibacillus catalpae sp nov., isolated from the rhizosphere soil of Catalpa speciosa. International Journal of Systematic and Evolutionary Microbiology 2013, 63, 1776-1781.
- Cheng, F.; Tang, C.; Yang, H.; Yu, H.; Chen, Y.; Shen, Z., Characterization of a Blend-Biosurfactant of Glycolipid and Lipopeptide Produced by Bacillus subtilis TU2 Isolated from Underground Oil-Extraction Wastewater. Journal of Microbiology and Biotechnology 2013, 23, (3), 390-396.
- Ma, Y.; Yu, H., Engineering of Rhodococcus cell catalysts for tolerance improvement by sigma factor mutation and active plasmid partition. Journal of Industrial Microbiology & Biotechnology 2012, 39, (10), 1421-1430.
- Jian, Z.; Yuchao, M.; Huimin, Y. J. I. J. S. E. M., Arthrobacter cupressi sp. nov., an actinomycete isolated from the rhizosphere soil of Cupressus sempervirens. 2012, 62, (Pt 11), 2731-2736.
- Jian, Z.; Ma, Y.; Yu, H. J. I. J. o. S.; Microbiology, E., Nocardioides lianchengensis sp. nov., a novel actinomycete isolated from soil of Liancheng county of Fujian province, China. 2012.
- Wang, Y.; Yu, H.; Zhang, J.; Luo, H.; Shen, Z., Double knockout of beta-lactamase and cephalosporin acetyl esterase genes from Escherichia coli reduces cephalosporin C decomposition. Journal of Bioscience and Bioengineering 2012, 113, (6), 737-741.
- Wang, Y.; Yu, H.; Song, W.; An, M.; Zhang, J.; Luo, H.; Shen, Z., Overexpression of synthesized cephalosporin C acylase containing mutations in the substrate transport tunnel. Journal of Bioscience and Bioengineering 2012, 113, (1), 36-41.
- Zhu, X.; Luo, H.; Chang, Y.; Su, H.; Li, Q.; Yu, H.; Shen, Z., Characteristic of immobilized cephalosporin C acylase and its application in one-step enzymatic conversion of cephalosporin C to 7-aminocephalosporanic acid. World Journal of Microbiology & Biotechnology 2011, 27, (4), 823-829.
- Sun, Y.; Yu, H.; Sun, X.; Shen, Z., Bi-steady state reaction kinetics of nitrile hydratase in free resting cells. Journal of Chemical Industry and Engineering (China) 2010, 61, (7), 1783-1789.
- Yu, H.; Ma, Y., Gene knockout strategies for metabolic pathway regulation in industrial microbes. Sheng wu gong cheng xue bao = Chinese journal of biotechnology 2010, 26, (9), 1199-208.
- Cheng, W. E. N.; Hui-min, Y. U.; Yun-peng, S. U. N.; Ying, W.; Zhong-yao, S., Efficient Quantification of Hyaluronic Acid in Fermentation Broth by Modified CTAB Method. Journal of Chinese Biotechnology 2010, 30, (2), 89-93.
- Luo, H.; Fan, L.; Chang, Y.; Ma, J.; Yu, H.; Shen, Z., Gene Cloning, Overexpression, and Characterization of the Nitrilase from Rhodococcus rhodochrous tg1-A6 in E. coli. Applied Biochemistry and Biotechnology 2010, 160, (2), 393-400.
- Ma, Y.; Yu, H.; Pan, W.; Liu, C.; Zhang, S.; Shen, Z., Identification of nitrile hydratase-producing Rhodococcus ruber TH and characterization of an amiE-negative mutant. Bioresource Technology 2010, 101, (1), 285-291.
- Sun, X.; Yu, H.; Shen, Z., Deactivation Kinetics of Nitrile Hydratase in Free Resting Cells. Chinese Journal of Chemical Engineering 2009, 17, (5), 822-828.
- Liu, C.; Yu, H.; Yuchao, M.; Pan, W.; Luo, H.; Shen, Z., Promoter recognition and beta-galactosidase reporter gene expression in Rhodococcus. Sheng wu gong cheng xue bao = Chinese journal of biotechnology 2009, 25, (9), 1360-5.
- Ma, X.-F.; Yu, H.-M.; Wen, C.; Luo, H.; Li, Q.; Shen, Z.-Y., Triple fusion of d-amino acid oxidase from Trigonopsis variabilis with polyhistidine and Vitreoscilla hemoglobin. World Journal of Microbiology & Biotechnology 2009, 25, (8), 1353-1361.
- Ling, T.; Zhu, J.; Yu, H.; Xie, L., Size Effect on Crystal Morphology of Faceted Face-Centered Cubic Fe Nanoparticles. Journal of Physical Chemistry C 2009, 113, (22), 9450-9453.
- Klein-Marcuschamer, D.; Santos, C. N. S.; Yu, H.; Stephanopoulos, G., Mutagenesis of the Bacterial RNA Polymerase Alpha Subunit for Improvement of Complex Phenotypes. Applied and Environmental Microbiology 2009, 75, (9), 2705-2711.
- Ling, T.; Xie, L.; Zhu, J.; Yu, H.; Ye, H.; Yu, R.; Cheng, Z.; Liu, L.; Liu, L.; Yang, G.; Cheng, Z.; Wang, Y.; Ma, X., Icosahedral Face-Centered Cubic Fe Nanoparticles: Facile Synthesis and Characterization with Aberration-Corrected TEM. Nano Letters 2009, 9, (4), 1572-1576.
Research
(1)Fundamentals of developing whole cell catalyst
Rhodococcus are organic-solvent tolerant strains with adaption abilities and diverse metabolic activities, thereby being widely utilized in bioconversion, biosynthesis and bioremediation. In recent years, the role of Rhodococcus ruber in biocatalysis has been increasingly highlighted. The advantageous features of R. ruber cells, such as high organic solvent tolerance and high target enzyme expression with low-cost urea inducer, have made them an ideal host for many exogenous enzymes with organic substrates. Thus, there is great potential to develop Rhodococcus cells for bioproduction of more interesting chemicals. In our lab, we on one hand focused on developing an efficient CRISPR/Cas9-mediated genome editing tool of R. ruber to overcome the limits of high GC-content (~70%) in genome, low transformation efficiency and recombination efficiency; on the other hand, we mainly worked on identifying the mechanism for high tolerance of R. ruber to the organic solvent (for example, the chaperone-networks in R. ruber and cell tolerance control via chaperones) and the mechanism for the unique urea-induced overexression of nitrile-degradation enzymes and the colony morphotype control from rough to smooth, etc.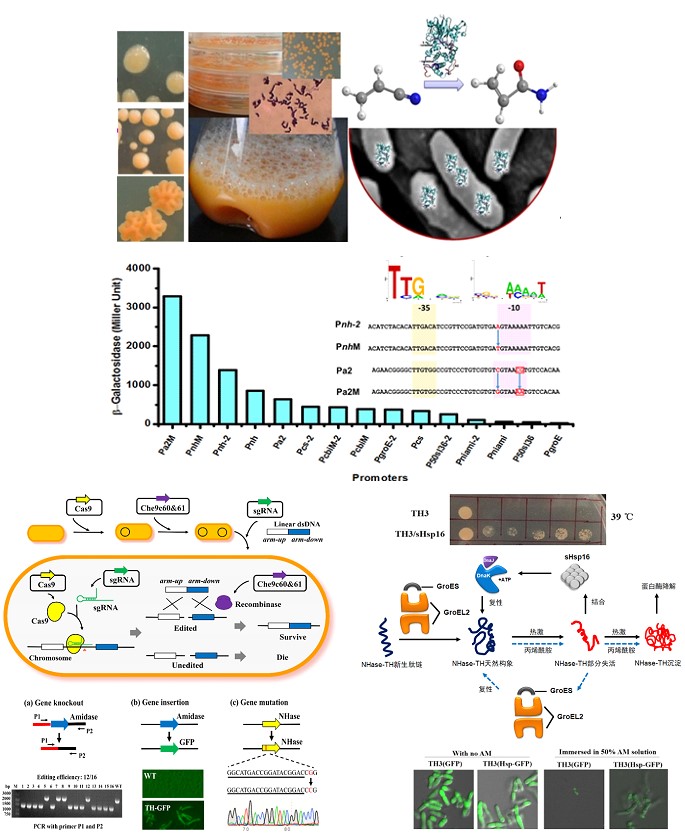
Rhodococcus cell biocatalysts, promoter engineering, Crispr/Cas9 genome editing tool and chaperones
(2)Enzyme engineering and whole cell catalysis
Through the coupling of salt bridge and disulfide bridge modified nitrile hydratase, recombinant Rhodococcus TH series with high tolerance, low/no by-products and other excellent properties were constructed. The strains above have been successfully applied in industry for many years, generating a great amount of profits, significant energy conservation and emission reduction. Both the economic and social benefits are remarkable. At present, we are working on Rhodococcus whole-cell biocatalyst as platform cells which can be widely used for the efficient catalytic synthesis of various amide, carboxylic acid and chiral pharmaceutical intermediates.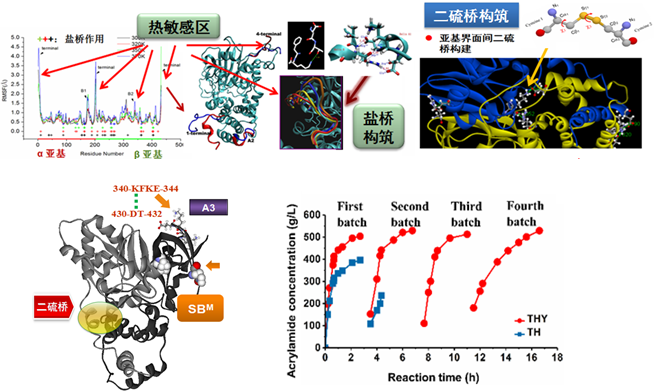
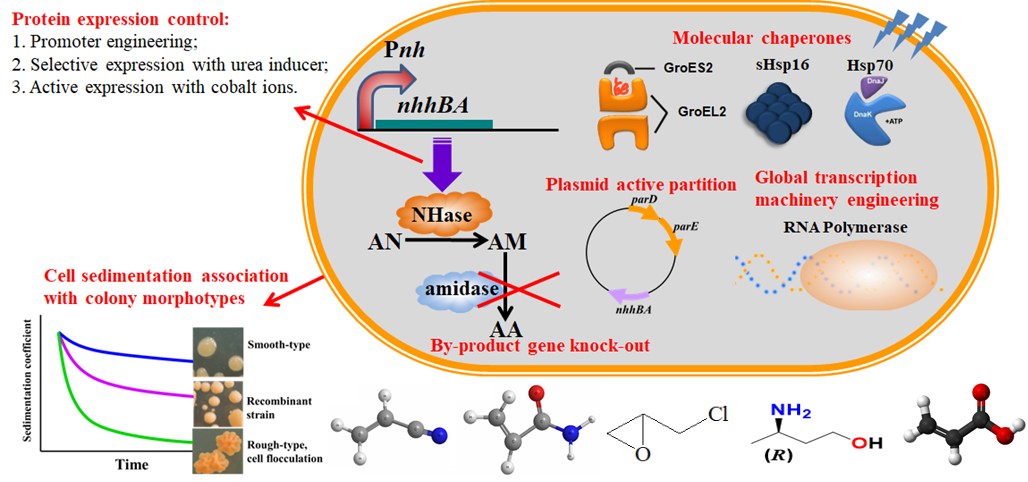
Designing salt-bridges and disulfide bonds in nitrile hydratase and the performance of the superior whole-cell Rhodococcus biocatalyst
(3)High-efficiency cell factory
Bacillus subtilis and Corynebacterium glutamicum have good biosafety and outstanding ability to synthesize target products efficiently. Our lab is dedicated to the creation of high-efficiency cell factories and metabolic engineering regulation in recombinant B. subtilis and recombinant C. glutamicum. The efficient synthesis of biomacromolecules were achieved such as lipopeptides (surfactants), hyaluronic acid and chondroitin.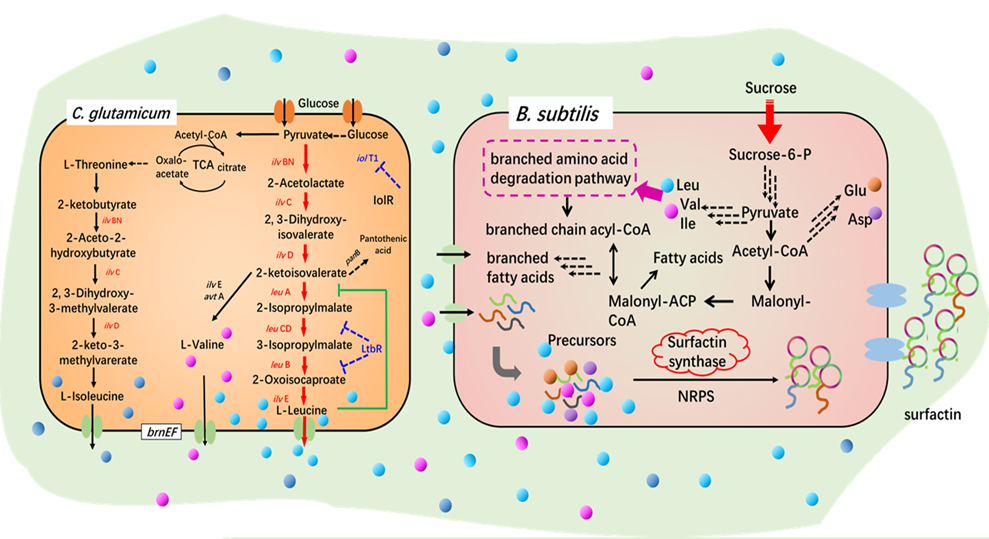
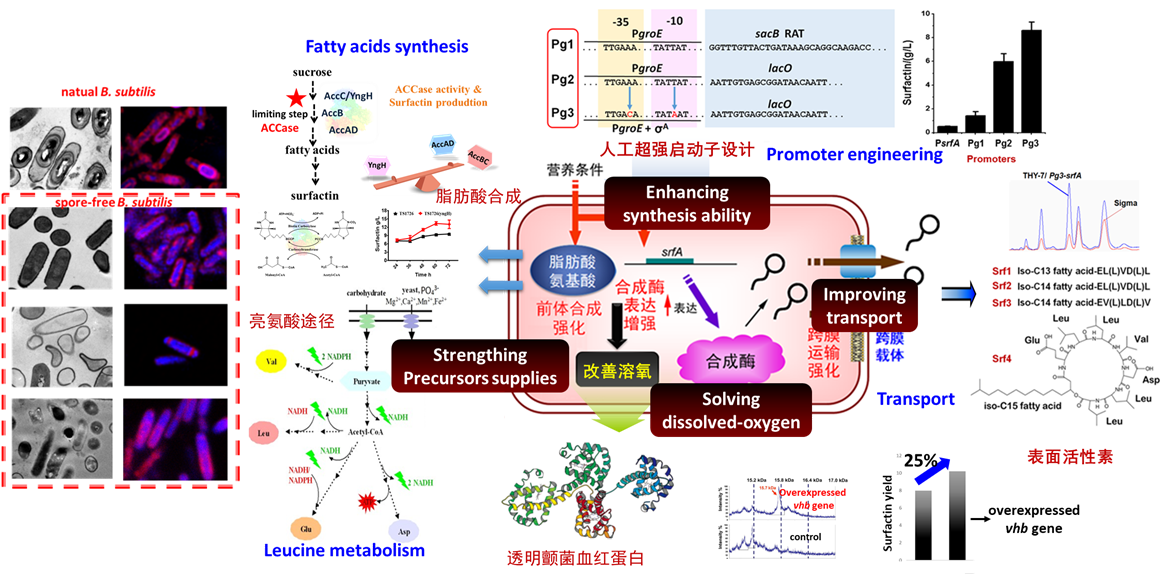
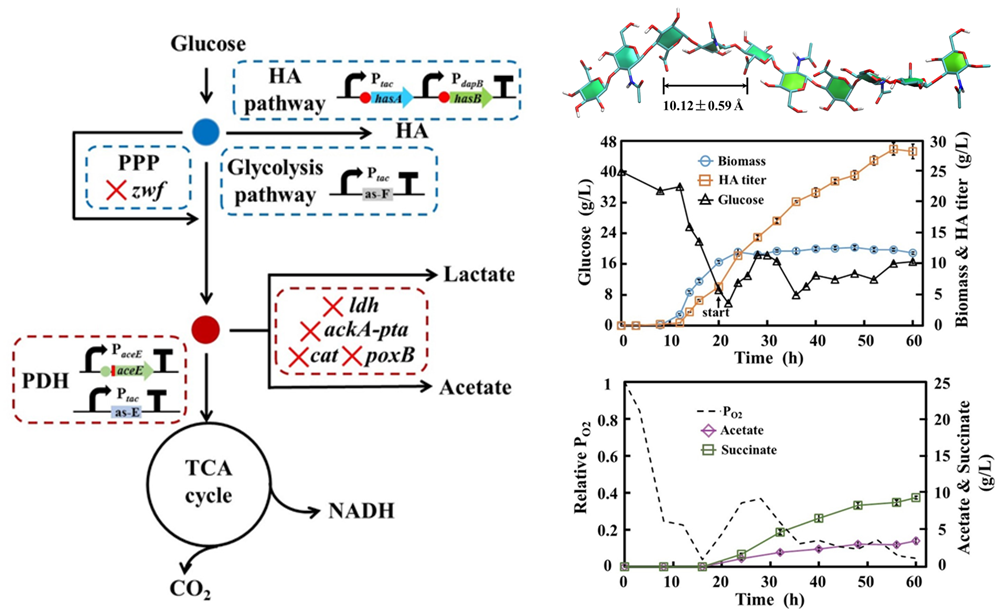
Bacillus subtilis and Corynebacterium glutamicum, surfactin and hyaluronic acid synthesis
(4)Nanobiotechnology
M13 bacteriophage is a member of filamentous bacteriophage family. Natural nano-scale size and special adsorption properties make M13 bacteriophage a hot research topic nowadays. Our group summarized the adsorption of metal ions by bacteriophages and prepared a variety of single metal nanoparticles and iron-nickel, iron-copper bimetallic nanoparticles using M13 bacteriophage as template. The Fe-Ni@M13 micro-electrolysis system was designed and constructed for the reduction of p-chloronitrobenzene. A new method of immobilization of enzymes mediated by M13 bacteriophage was established, which also provides new ideas for the further studies.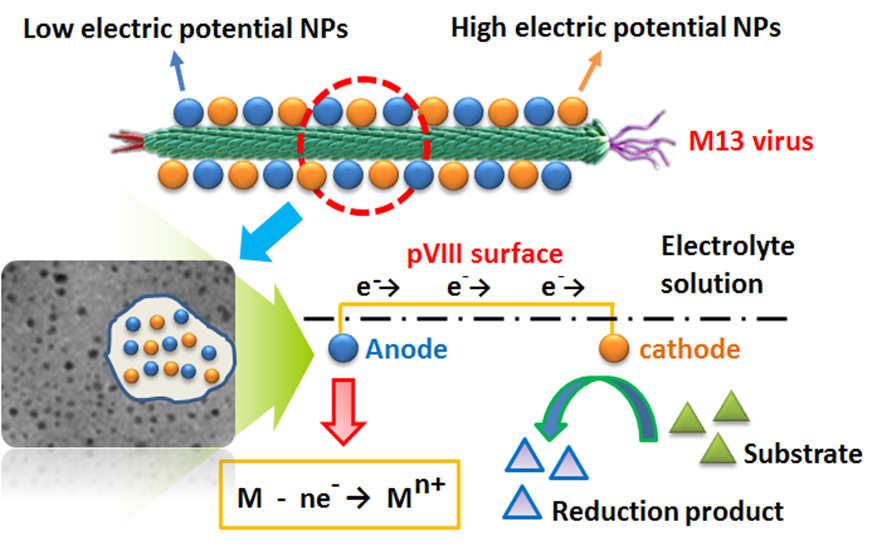
Fe-Ni@M13 micro-electrolysis system
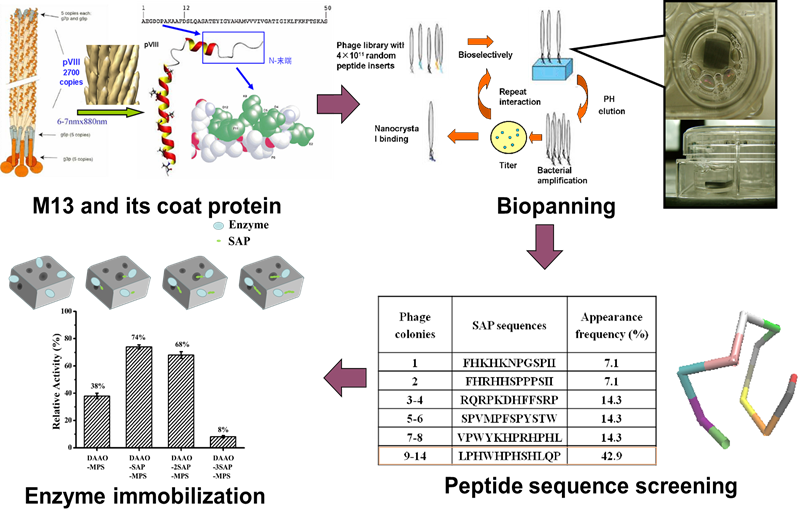
A novel immobilized enzyme method based on M13 phage



